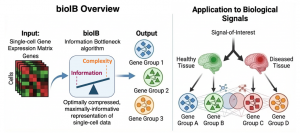
Modern technologies can now measure the activity of thousands of genes in individual cells, offering unprecedented insight into how cells function in health and disease. However, this wealth of data is often so complex that it becomes difficult to extract specific biological signals, such as disease progression.
To address this challenge, we developed bioIB, a computational method based on the Information Bottleneck algorithm*, that seeks to extract from single-cell data the most relevant information to a particular biological signal or process. Given a signal of interest – for example, annotations of cells to different types or originating from healthy vs. diseased tissues – bioIB generates a compact, interpretable representation of the data, highlighting only the meaningful biological differences related to the signal of interest. Beyond simplifying the data, bioIB also reveals how these gene groups relate to one another, uncovering connections between biological processes and cell types.
bioIB can be applied across a wide range of biological systems, including neurodegeneration, development, and epithelial-to-mesenchymal transition, revealing interpretable representations of multicellular processes and transitions underlying dynamical processes across time scales. In particular, we used bioIB to identify gene groups involved in the transition from healthy brain cells to those affected by Alzheimer’s disease.
Significance
BioIB provides a powerful way to extract meaningful insights from complex biological data. Producing compact, signal-specific, and interpretable representations helps researchers focus on the biological processes most relevant to their questions. BioIB can thus be used to accelerate the discovery of disease mechanisms, improve the identification of potential biomarkers, and deepen our understanding of how cellular behavior changes in health and disease.
—————————————————-
* We, at ELSC and the School of Computer Science and Engineering, are proud of this collaboration, which continues the legacy of the late Prof. Naftali (Tali) Tishby, who introduced the Information Bottleneck framework in 1999. This theory provides a principled way to balance accuracy and complexity by retaining only the most relevant information while discarding the rest. It has also inspired the idea that the brain may operate in a similar way, actively compressing sensory input to focus on what matters most. More recently, in 2023, it was mathematically shown that controlling the Information Bottleneck can help control generalization error in deep learning, highlighting its fundamental importance in both neuroscience and artificial intelligence.
References
Tishby, N., Pereira, F. C., & Bialek, W. (1999). The information bottleneck method. The 37th annual Allerton Conference on Communication, Control, and Computing.
Kawaguchi, K., Deng, Z., Ji, X., Huang, J. (2023) How does information bottleneck help deep learning? International conference on machine learning; PMLR.
Illustration (Generated using FigureLabs)